
17/4/2026
•
Kimialys
•
4 min

This application note builds on work performed by Kimialys for the client Tollys, as part of a study published in Microbial Cell in June 2023.
RNA is well known for being difficult to handle due to its low stability and rapid degradation by RNases, even at room temperature. Consequently, immobilizing RNA oligomers on a biosensor surface and monitoring nucleoprotein interactions require strict precautions, including maintaining RNase-free conditions.
K-One is Kimialys’ proprietary technology platform, representing the culmination of over 10 years of academic research conducted at Paris-Saclay University. It stands as a flexible surface treatment that can be easily adaptable to various applications with a full control over ligand distribution and accessibility as well as elimination of non-specific interactions.
In conclusion, K-One delivers robust, sensitive, and reproducible measurements, while optimizing the Signal to Noise Ratio (SNR), even under challenging conditions such as detecting target molecules diluted in complex media (e.g., human serum or cell culture supernatant) or working with fragile ligands (RNA oligomers).
This application note demonstrates the efficacy of Kimialys' K-One surface chemistry in monitoring nucleoprotein interactions between immobilized RNA oligomers and a target protein using SPRi technology.
Surface Plasmon Resonance imaging (SPRi) is a sensitive, reliable, and fast optical biosensing technique widely used to monitor biological interactions in real-time. It utilizes a gold biochip surface functionalized with ligands that specifically interact with target molecules in a solution. The dynamics of complex association and dissociation at the biochip surface allow for the characterization of association and dissociation rates constants and of the complex affinity. Additionally, multiplexing capability of the SPRi technology enables multiple, parallel measurements, allowing for simultaneous analysis of different interactions.
Example of one injection cycle is presented in Figure 1.

SPRi sensorgrams of one experimental cycle: injection of 36 nM target protein (association phase, t = 0 s to t = 180 s), analyte dissociation (t = 200 s to t = 1400 s), and two surface regeneration injections of 30 s each (at t = 1400 s). The entire process was recorded on selected spots of the biosensor surface where RNA and DNA were covalently immobilized at different concentrations. "Background" refers to the signal registered directly on the K-One biochip surface.
Analysis of the obtained data confirmed specific interactions between the RNA oligomer and the injected target protein. The intensity of the registered signal correlated with the surface density of the grafted RNA. The analyte did not interact with the control DNA or the background (the biosensor surface where no ligand was present), indicating that only specific interactions were detected. After the dissociation phase, the entire surface of the biosensor was completely regenerated (Figure 1, t = 1600 s), allowing for multiple injection/regeneration cycles.
Three different RNA surface densities were tested to determine the conditions where affinity constant could be extracted with 1:1 binding model. RNA ligands were spotted on a single biochip in duplicate at the concentrations 10, 20 and 50 µM. It allowed to measure nucleoprotein interactions in 6 replicates. Sensorgrams collected during the injections of the target protein at increasing concentrations were used to determine the affinity (KD) of the RNA-protein complex. Background signals were subtracted from the signals recorded at the RNA spots to ensure accuracy of the measurements. The maximum response, measured at equilibrium (at the end of the association step, t = 180 s), was plotted as a Langmuir binding isotherm and fitted to determine KD, as shown in Figure 2. To confirm repeatability, measurements were performed on three different biochips.

Superimposed sensorgrams showing injections of the target protein at concentrations ranging from 0 to 36 nM over a spot of 20 µM RNA.
.png)
Langmuir binding isotherm plotting signal intensity at t = 180 s versus the concentration of the injected target protein. Experimental data were fit to the Langmuir model.
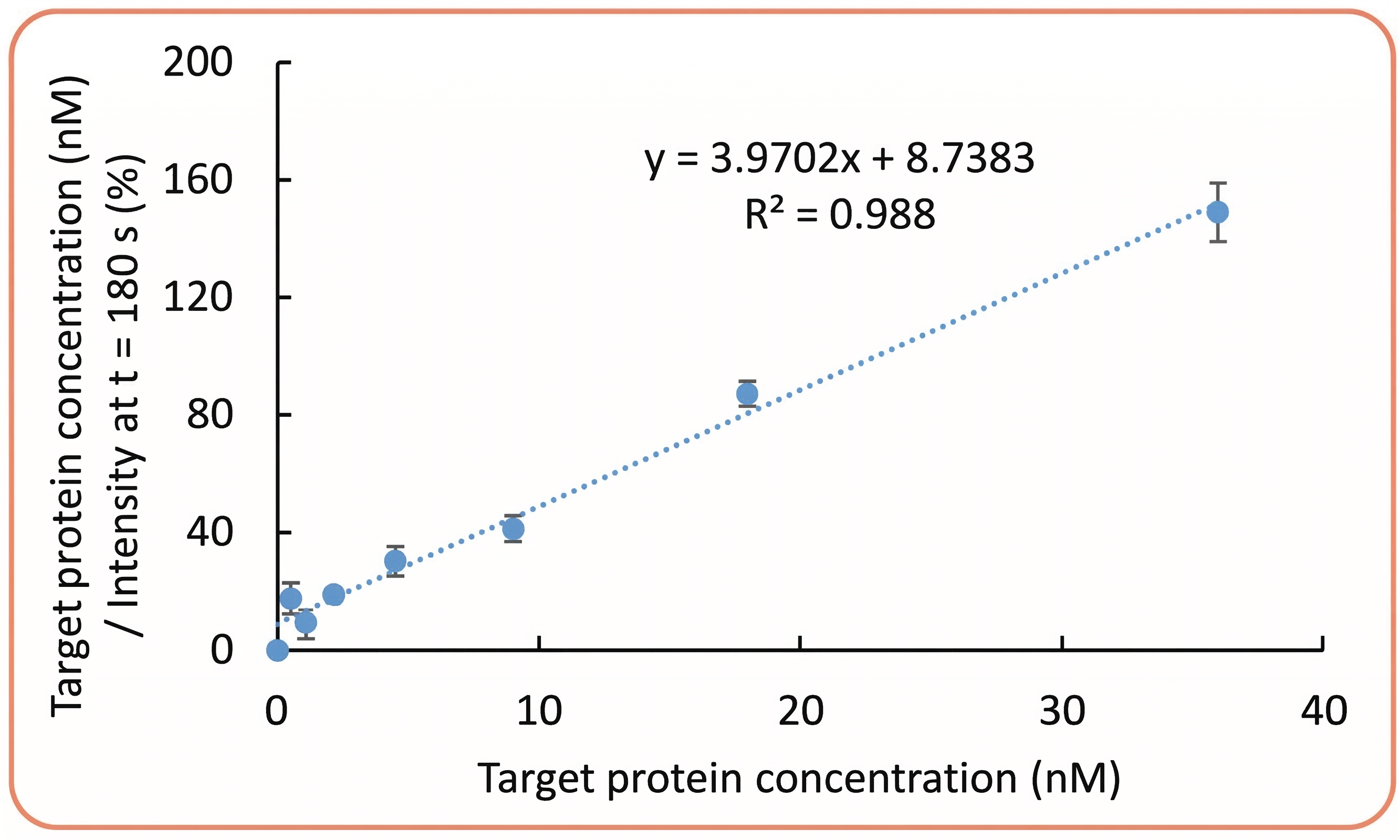
Linearization of the Langmuir binding isotherm.
Error bars on Figures B and C correspond to the standard deviation of two measurements collected on two different spots of 20 µM RNA.
The KD calculated from three independent experiments was 1.94 ± 0.20 nM. Kinetic constants were also extracted by fitting the sensorgrams using binding 1:1 model with BIAevaluation software. While the association phase was challenging to fit, the fits of the dissociation phases were of a good quality. Such-calculated dissociation was of (2.31 ± 0.40) · 10-4 1/s. Considering that the previously obtained KD was 1.94 ± 0.20 nM, one may deduce the association rate to be at 7.60 · 104 1/Ms. In addition, the three different ligand densities evaluated in this study did not influence the obtained ketic constants and affinity.
In the presented study, the RNA oligomers immobilized on K-One biosensor surface were stable, enabling accurate measurement of their interactions with the target protein. Importantly, no non-specific interactions were detected, either with the negative control (DNA) nor with the biosensor surface (background). The data obtained from these interactions were used to extract kinetic constants and determine the KD value. The experiments were repeated independently three times, demonstrating good reproducibility of the results.
Throughout this study, we have demonstrated the potential of K-One surface chemistry in handling fragile ligands, such as RNA oligomers, while delivering high-quality results and affinity measurements. Moreover, Kimialys is not just about K-One technology — first and foremost, we are a team of biosensor engineers and chemists with strong expertise in Surface Plasmon Resonance assay development, making Kimialys your partner of choice for the most challenging SPR analyses, providing a level of expertise and support that stands out in the field.
Kimialys is an innovative industrial company specialized in biosensing, enabling laboratories worldwide to develop and produce tomorrow’s diagnostics and therapeutics. With over 100 projects delivered on complex real-world applications, we are contributing to the most innovative assays under development.
K-One is Kimialys’ proprietary technology, a surface chemistry that powers all our products and services today. It is a self-assembled monolayer (SAM) optimized for gold nanoparticles, chips, and electrodes unlocking applications once out of reach due to technological limitations and setting new standards for surface treatment in biosensing (detection, extraction, screening, characterization).
By combining K-One with our expertise in ligand functionalization, nanoparticle-based assays, and biosensors, we enable advanced applications in diagnostic development, drug discovery, and bioanalysis. Our approach enhances key assay parameters, such as sensitivity, specificity, selectivity, reproducibility, without ever compromising one for another, as well as cost-effectiveness.

Subscribe to our monthly newsletter and stay informed about the latest news, application notes, and insights from our activities.
By clicking Sign Up you're confirming that you agree with our Terms and Conditions.